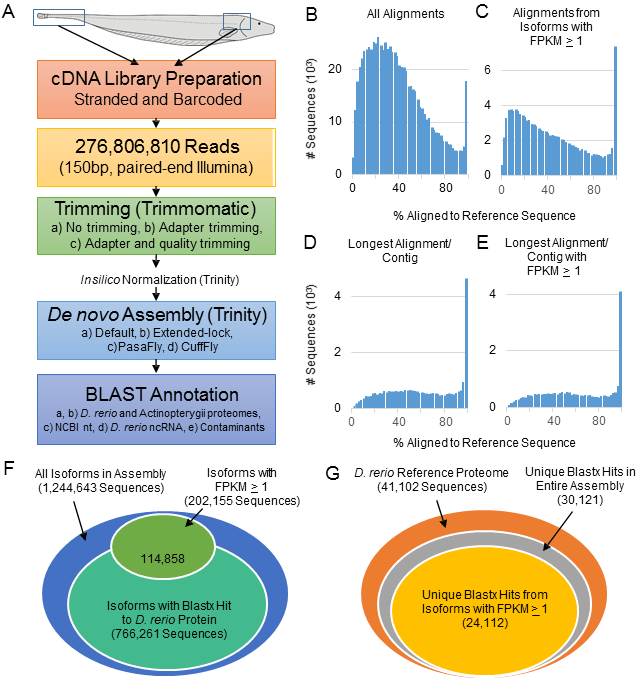
Transcriptome assembly workflow and alignment to D. rerio reference proteome. A. RNA was extracted from brain and spinal cord tissue of adult A. leptorhynchus, fragmented and barcoded, and strand-specific cDNA libraries were created for Illumina sequencing. Reads were trimmed using several strategies and then normalized in silico prior to de novo assembly with Trinity. Transcript reconstruction was performed using several strategies and then benchmarked using BLAST to maximize transcripts aligning to a D. rerio reference proteome. The best assembly was then further annotated. B. Out of the transcripts from the entire A. leptorhynchus assembly with any alignment to a D. rerio reference protein, most transcripts aligned to <40% of the reference D. rerio proteins. C. When filtering the assembly for transcripts with FPKM ≥ 1, the relative proportion of transcripts with less than complete alignments was reduced. D. When selecting only the best alignment for a given D. rerio reference protein, this best alignment was more complete than many other fragmented alignments, which can be partially attributed to multiple transcripts from the same gene with less than complete ORFs. E. The distribution of the longest aligned sequences for each unique D. rerio reference sequence was preserved in the assembly containing only transcripts with FPKM ≥ 1. F. Out of the entire assembly, more than half of the transcripts had at least some alignment to a reference D. rerio sequence. Similarly, more than half of the sequences with FPKM ≥ 1 aligned to a D. rerio protein sequence. G. Using only transcripts with FPKM ≥ 1, nearly 60% of the reference D. rerio proteome had at least one assembly transcript with an alignment (24,112 sequences), which represented approximately 80% of the D. rerio sequences that were hit by the entire assembly (30,121 unique sequences). (From Salisbury et al., 2015.)
Publication:
Salisbury, J.P., Sîrbulescu, R.F., Moran, B.M., Auclair, J.R., Zupanc, G.K.H., Agar, J.N.: The central nervous system transcriptome of the weakly electric brown ghost knifefish (Apteronotus leptorhynchus): de novo assembly, annotation, and proteomics validation. BMC Genomics 16, 166 (2015), doi:10.1186/s12864-015-1354-2




